The IPT and MDT
Overview
Teaching: 60 min
Exercises: 0 minQuestions
What are the IPT and MDT?
How do you use them to publish metabarcoding data?
Objectives
Be able to explain what the IPT and MDT are
Know how to find and use guidance for the IPT and MDT
The Integrated Publishing Toolkit (IPT)
- User Guide: https://ipt.gbif.org/manual/en/ipt/latest/
- DNA Derived Data Publishing Guide: https://doi.org/10.35035/doc-vf1a-nr22
What is the IPT?
The Integrated Publishing Toolkit (IPT) is an open source web application that helps users create and publish Darwin Core Archives (DwC-A) to GBIF. In the context of publishing DNA-derived data, this means that ayn occurrence data that includes information relevant to the DNA-derived data extension can be uploaded, mapped to Darwin Core, and published to GBIF.
Yes!
The IPT can handle DNA barcoding, metabarcoding datasets, metagenomic, qPCR and other data types. But it may require some data wrangling to align the data with the Darin Core and EML standards.
No!
The IPT cannot handle: raw sequencing data (e.g Fastq files), OTU tables in a community matrix format or FASTA files. This is because it is a general tool for producing a DwC-A. The MDT (described below) is more appropriate for wrangling and publishing typical outputs from metabarcoding studies.
How does it work?
You can think of the IPT as a program, like MS Excel. Just like Excel produces spreadsheets and workbooks, the IPT produces Darwin Core Archives and allows you to register (i.e. publish) that file with GBIF and other networks. A typical workflow includes:
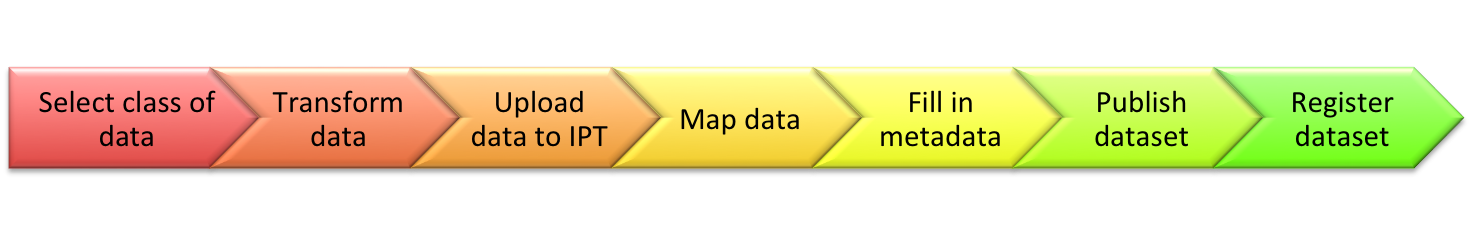
Transforming and mapping your DNA data to Darwin Core is best guided by:
Abarenkov K, Andersson AF, Bissett A, Finstad AG, Fossøy F, Grosjean M, Hope M, Jeppesen TS, Kõljalg U, Lundin D, Nilsson RN, Prager M, Provoost P, Schigel D, Suominen S, Svenningsen C & Frøslev TG (2023) Publishing DNA-derived data through biodiversity data platforms, v1.3. Copenhagen: GBIF Secretariat. https://doi.org/10.35035/doc-vf1a-nr22.
There are a variety of tools you might use to wrangle your data, including OpenRefine, R, and Python.
Example Dataset
We will demo the IPT with the same example dataset we provided in the previous example, however we have already taken the time to split it into a table of occurrence terms and one of DNA terms. It is a slightly modified verion of a real dataset with COI metabarcoding of DNA extracted from sea water. The dataset has rich metadata and is a good example of a well-documented dataset. This data was originally published as:
Shea M M, Boehm A B (2024). COI data from: Environmental DNA metabarcoding differentiates between micro-habitats within the rocky intertidal (Shea & Boehm, 2024). Version 1.5. United States Geological Survey. Occurrence dataset. https://ipt-obis.gbif.us/resource?r=shea_boehm_2024&v=1.5 https://doi.org/10.15468/33artc accessed via GBIF.org on 2025-05-07.
You can download the files here:
- https://sunray1.github.io/2025-05-09-GBIF-NA-DNAPublishing/files/DNA_example_2_for_IPT_occ.csv.zip
- https://sunray1.github.io/2025-05-09-GBIF-NA-DNAPublishing/files/DNA_example_2_for_IPT_dna.csv.zip
The Metabarcoding Data Toolkit (MDT)
- User Guide: https://docs.gbif-uat.org/mdt-user-guide/en/
- GBIF-US MDT: https://mdt.gbif.us/
- MDT Homepage: https://mdt.gbif.org/

What is the MDT?
The Metabarcoding Data Toolkit (MDT) is an open source web application developed as part of a pilot phase of the GBIF Metabarcoding Data Programme. It helps users publish DNA-derived data to GBIF. The MDTs can be used by anybody. GBIF nodes who wish to administer an instance of the MDT can apply for a GBIF hosted MDT.
Yes!
The Metabarcoding Data Toolkit can handle DNA metabarcoding datasets (aka amplicon sequence data) – specifically OTU tables and their associated metadata.
No!
The MDT cannot handle: raw sequencing data (e.g Fastq files), metagenomic/shotgun datasets, specimen barcodes, qPCR
How does it work?
You can think of the MDT as something like an IPT that is expecting typical outputs from processed metabarcoding (AKA eDNA) data. As illustrated below, this typically, this will include an OTU table, a taxonomy table, a samples table, representative OTU sequences in FASTA format, and study level metadata.
The MDT is designed to take import that data and help users (1) map it to Darwin Core, (2) create EML metadata, and (3) publish the dataset with GBIF. It has more features that we will cover in the demo.
 Figure 8 from MDT User Guide [link]: An example OTU_table with OTU IDs linked to the Taxonomy table and Sample IDs linke to the Samples table. OTU_table: sequence read counts of each OTU per sample; Taxonomy: DNA sequences and taxonomy per OTU; Samples Sample metadata per sample; Study (optional): Metadata values applying to all samples and OTUs; Seqs.fasta (optional) OTU sequences in fasta format.
Figure 8 from MDT User Guide [link]: An example OTU_table with OTU IDs linked to the Taxonomy table and Sample IDs linke to the Samples table. OTU_table: sequence read counts of each OTU per sample; Taxonomy: DNA sequences and taxonomy per OTU; Samples Sample metadata per sample; Study (optional): Metadata values applying to all samples and OTUs; Seqs.fasta (optional) OTU sequences in fasta format.
🤓 Nerd Bonus! MDT has an API 🤓
We’re not going to go over it today, but for those who are interested in making programmatic connections to the MDT, it does have an API. Documentation can be found in the user guide: https://docs.gbif-uat.org/mdt-user-guide/en/index.en.html#using-mdt-through-api
There aren’t many examples of leveraging the API yet, but it’s an exciting capability that sets it apart from the IPT.
Example Dataset
- MDT Sandbox: https://mdt.gbif-test.org/
- MDT User Guide Example Dataset 2: https://docs.gbif-uat.org/mdt-user-guide/en/index.en.html#example2

We will use the example dataset provided in the MDT User Guide to demo the MDT. It is a real dataset with COI metabarcoding of DNA extracted from sea water. The dataset has rich metadata and is a good example of a well-documented dataset. This data was originally published as:
Shea M M, Boehm A B (2024). COI data from: Environmental DNA metabarcoding differentiates between micro-habitats within the rocky intertidal (Shea & Boehm, 2024). Version 1.5. United States Geological Survey. Occurrence dataset. https://ipt-obis.gbif.us/resource?r=shea_boehm_2024&v=1.5 https://doi.org/10.15468/33artc accessed via GBIF.org on 2025-05-07.
The example version has been modified slightly from the original dataset.
- Download file: Example Dataset 2.
https://docs.gbif-uat.org/mdt-user-guide/example_data/example_data2current.en.xlsx
Demo
We’re doing a live demo, but this recording of a previous demo by GBIF will be here for future reference:
Key Points
The IPT is a tool for turning data into a Darwin Core Archive. You can include the DNA derived data extension in the Darwin Core Archive.
The IPT has an excellent manual that can be used in combination with the DNA Publishing Guide for self-teaching.
The MDT is a tool for turning typical metabarcoding outputs into a Darwin Core Archive.
The MDT is available through a GBIF pilot program.
There is a great user guide and example datasets to help you learn how to use the MDT.