Dataset Mapping
Overview
Teaching: 30 min
Exercises: 60 minQuestions
How do I map a DNA dataset to Darwin Core?
Objectives
Become familiar with exploring DNA data and mapping it to Darwin Core terms.
If you brought a dataset, begin to map it to Darwin Core.
Introduction
We’re going to map a barcoding dataset as a group, and then have time to work individually on data your brought with you, or an example metabarcoding dataset. First let’s make sure we understand the difference between enriched occurrences (e.g. barcoding), DNA-derived occurrences (e.g. metabarcoding), and targeted species detection (e.g. qPCR). Sections 2.2.1 and 2.2.2 of the guide has a table with recommendations for mapping both types to Darwin Core and the DNA extension.
Enriched Occurrence (barcoding)
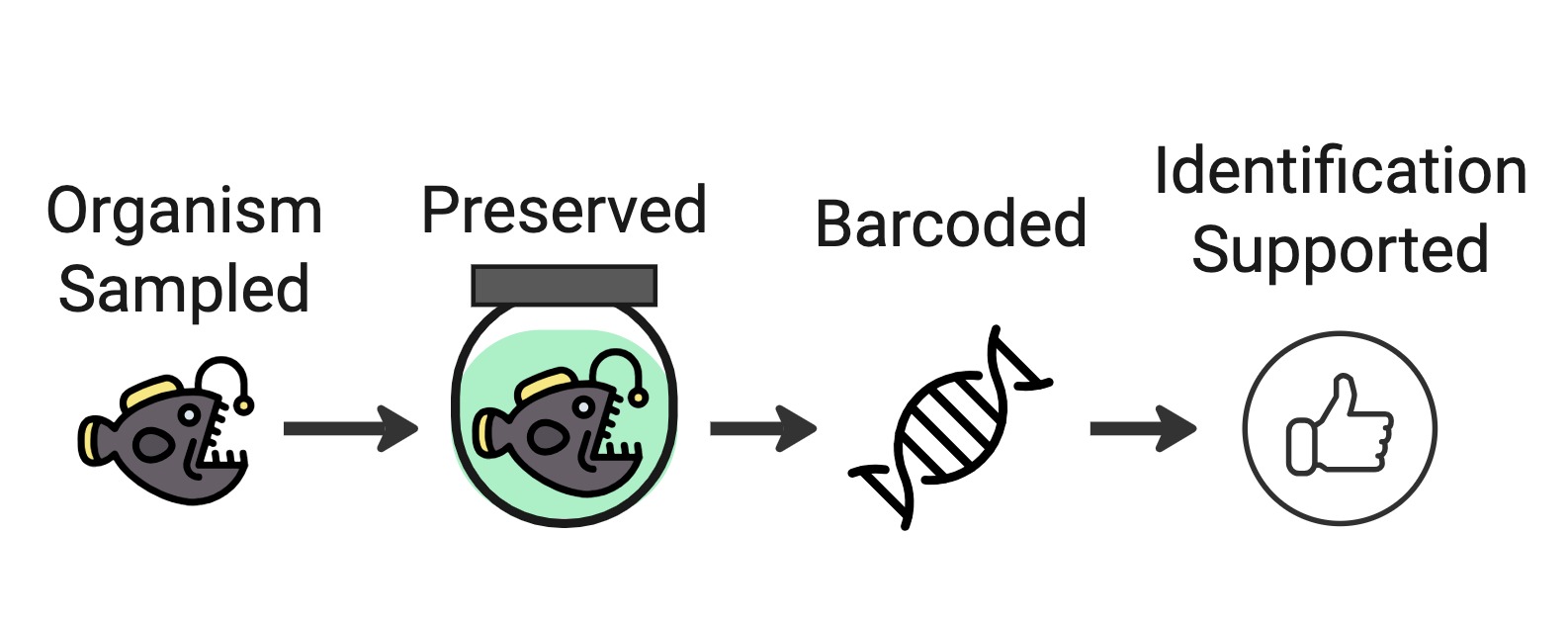
In the GBIF-OBIS guide to DNA publishing, this is described as ‘Category II’:
2.1.2. Category II: Enriched occurrences If some genetic material is, or can be, associated with an observation or a specimen, we will categorize this type of data as “enriched occurrences”. In this context, the sequences are not the only evidence of occurrences. One can always trace the information back to a vouchered specimen or observed organism. This category includes barcoding datasets and some DNA metabarcoding datasets with reference material for example. For more guidance on barcoding, follow Centre for Biodiversity Genomics, University of Guelph (2021).
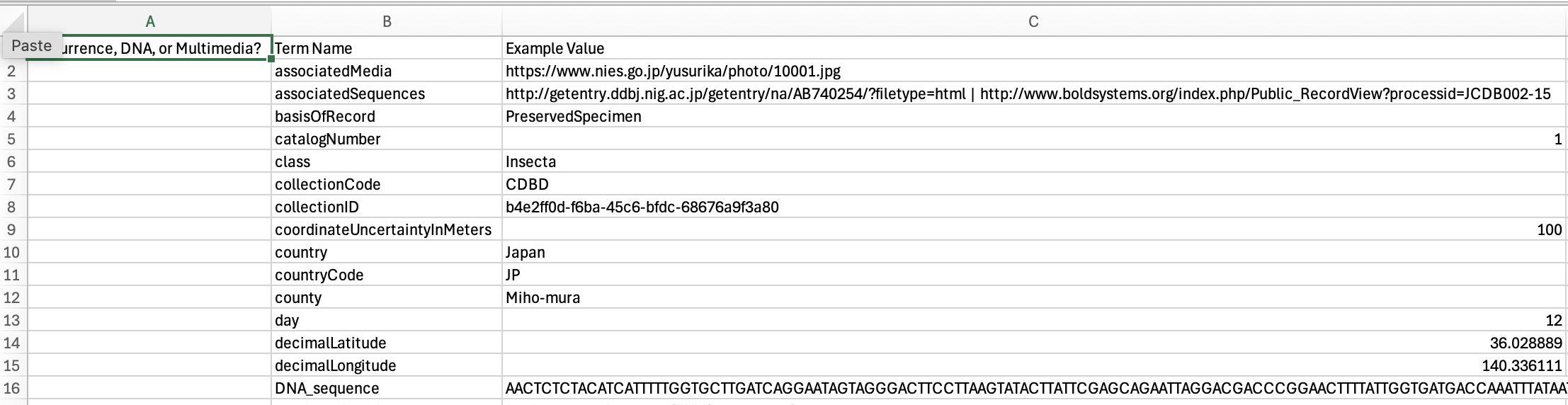
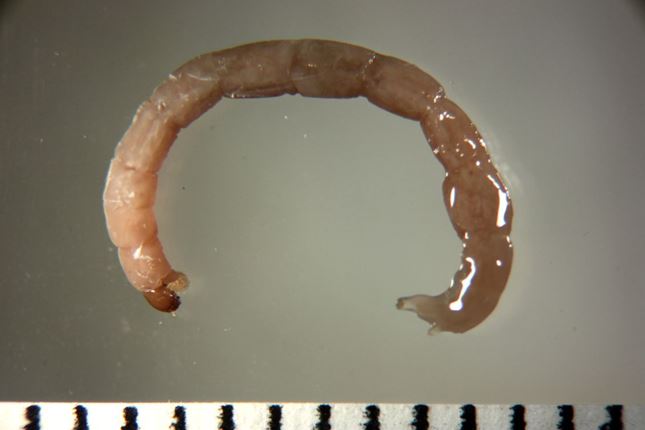
Explore this occurrence. What information is different, or extra, compared to a typical museum specimen published through GBIF?
Here is an example enriched occurrence from the dataset, Chironomid Specimen records in the Chironomid DNA Barcode Database.
DNA-derived occurrences (e.g. metabarcoding AKA eDNA)
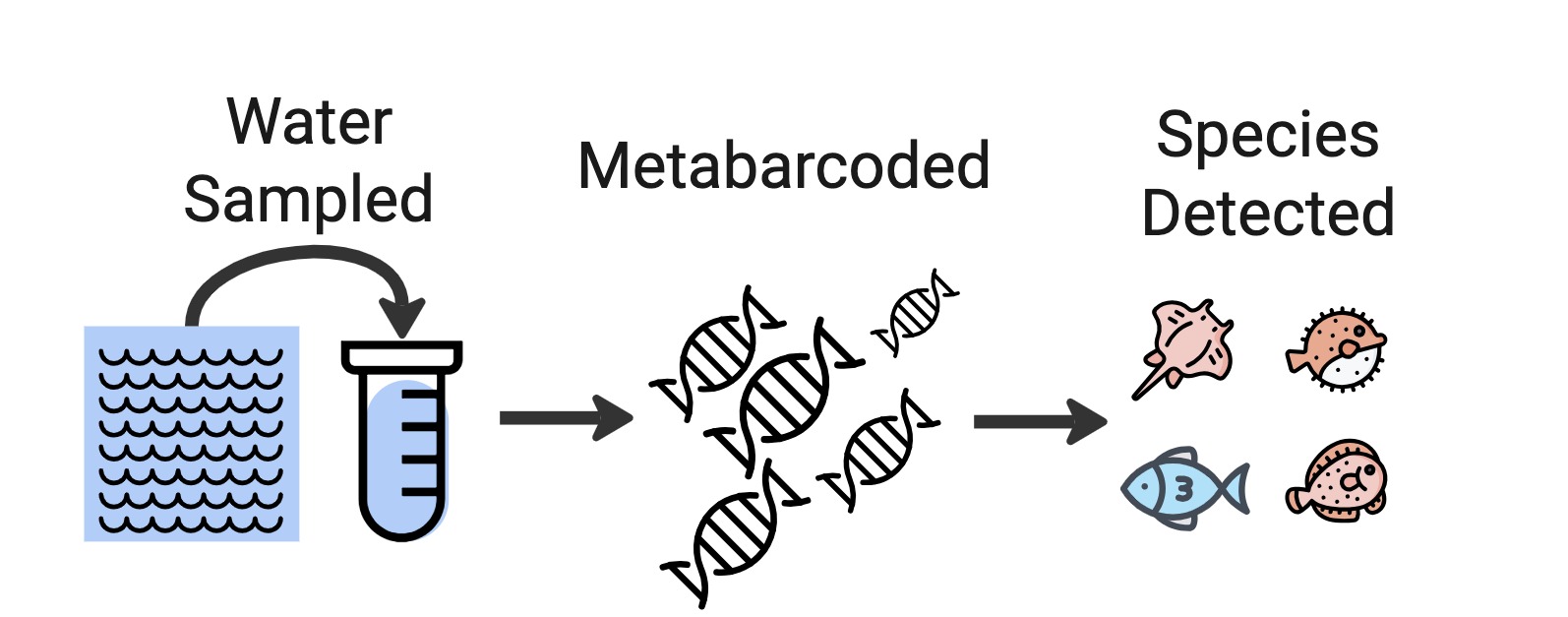
In the GBIF-OBIS guide to DNA publishing, this is described as ‘Category I’:
2.1.1. Category I: DNA-derived occurrences This category concerns data where a DNA sequence or detection through PCR is the only evidence for the presence of a given organism or community. In other words, the data cannot be traced back to an observable specimen. This is the case for many metagenomics, metabarcoding and eDNA studies.
Depending on what you’re used to, this may seem quite different from other biological data types.
Explore this occurrence. What information is different, or extra, compared to the enriched occurrence you explored above?
Here is an example DNA-derived occurrence from the dataset, COI data from: Environmental DNA metabarcoding differentiates between micro-habitats within the rocky intertidal (Shea & Boehm, 2024).
Targeted Species Detection (e.g. qPCR)
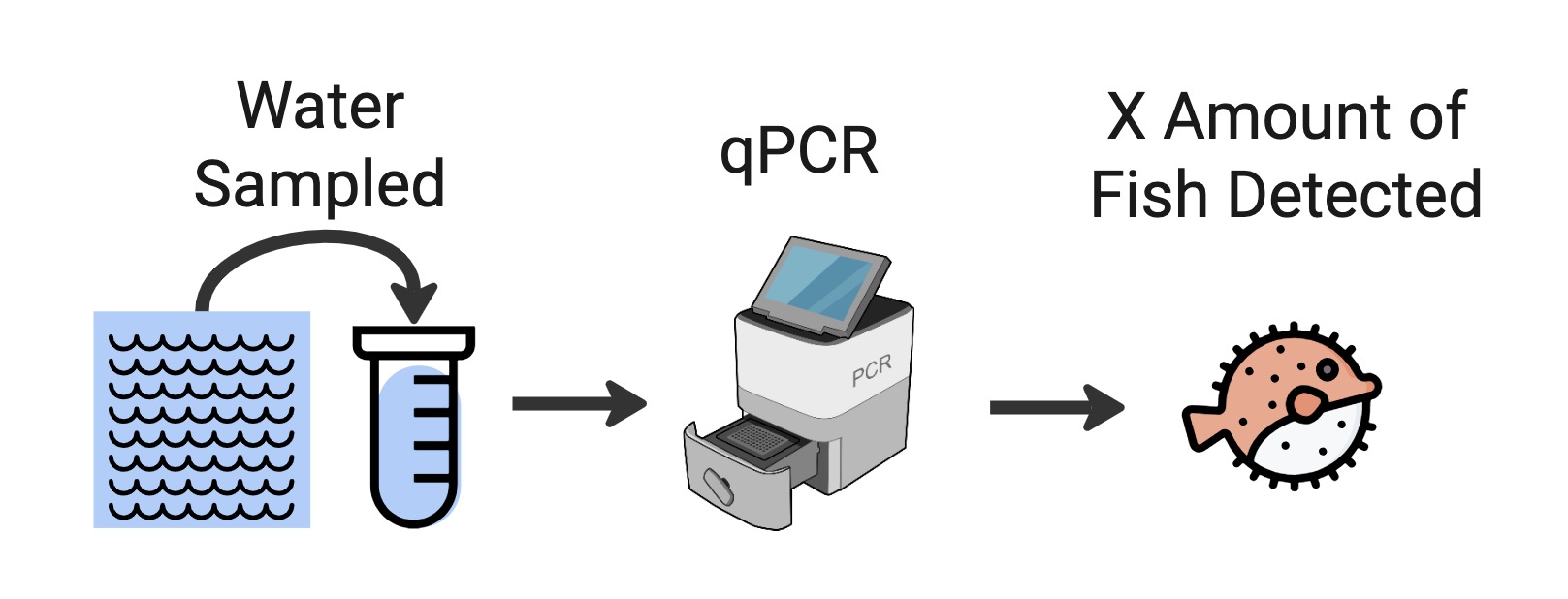
In the GBIF-OBIS guide to DNA publishing, this is described as ‘Category III’:
2.1.3. Category III: Targeted species detection (qPCR/ddPCR) This category concerns data where a specific (qPCR/ddPCR) assay is used to detect the presence (or absence) of a DNA sequence specific to the target organism in an environmental sample. In this case the occurrence record may not even contain sequence data, as it is the process itself that determines the occurrence. With qPCR/ddPCR analyses for targeted species detection, many studies also report absence of that specific species for a given sample. Absence data is highly dependent on the detection limit of the specific assay, as well as field and lab protocols. As for DNA-metabarcoding data there is an issue of both false negatives and false positives, and it is important that sufficient information is reported for evaluating the records.
Depending on what you’re used to, this may seem quite different from other biological data types.
Explore this occurrence. What information is different, or extra, compared to the enriched occurrence you explored above?
Here is an example targetted species detection occurrence from the dataset, Kenai National Wildlife Refuge Invasive Fish Surveys - 2023.
How to structure your Darwin Core Archive?
The structure of your data and/or experiment will help you decide the best structure for publishing (e.g. which core to use, which extensions to use, how many tables do you need?). In case you’re not very familiar with Darwin Core Archives, we review possible structures here.
Occurrence Core
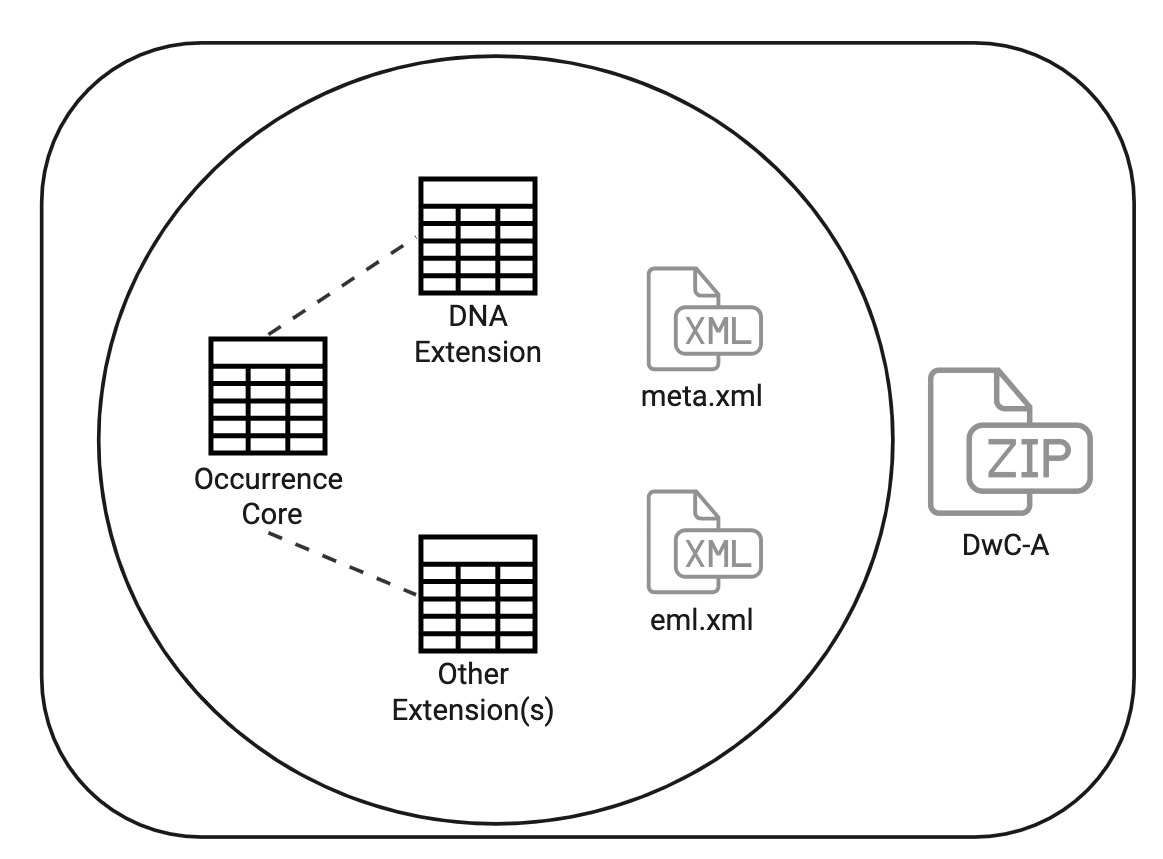
Until recently, Darwin Core Archives (DwC-A) for DNA data had to be structured as an occurrence table and a DNA table. Like any DwC-A, this would include two XML metadata files. Extensions that can be linked to the occurrence core table could also be used.
Event Core
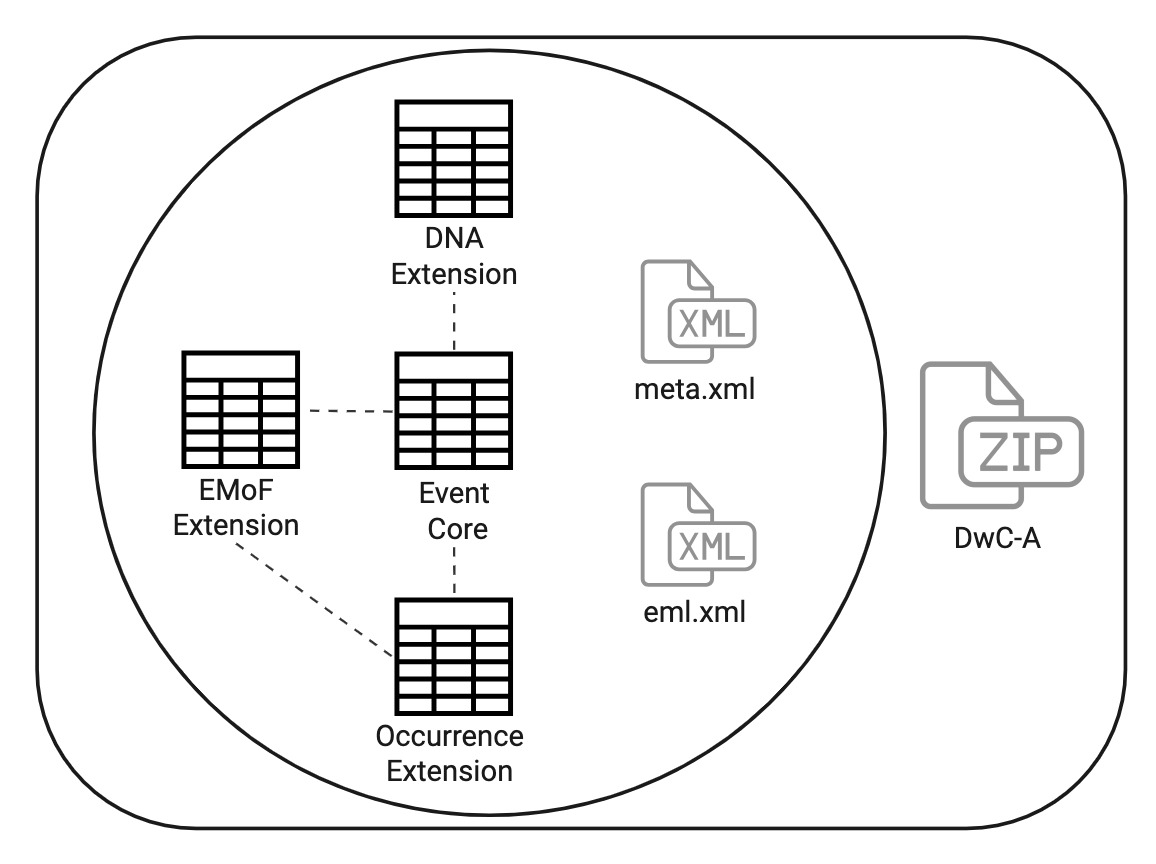
Last year, a tweak was made to to the extension to allow Event Core and relevant extensions to be used. However, for today we'll be keeping it simple and only working with an occurrence table and DNA table.
Group Mapping exercise
Dataset: Chironomid Specimen records in the Chironomid DNA Barcode Database https://doi.org/10.15468/hxhow5
Review these terms an example values from the dataset. Note which terms should go in the occurrence table, the DNA table, and the multimedia table.
Download this file: https://sunray1.github.io/2025-05-09-GBIF-NA-DNAPublishing/files/enriched_occ_mapping_practice.csv
Don’t forget to lean on your guidance!
Individual Mapping Excercise
Try mapping your data to Darwin Core and the DNA Extension. We're hear to discuss and help!
Use the example metabarcoding dataset from the MDT User Guide. It is a slightly modified verion of a real dataset with COI metabarcoding of DNA extracted from sea water. The dataset has rich metadata and is a good example of a well-documented dataset. This data was originally published as:
Shea M M, Boehm A B (2024). COI data from: Environmental DNA metabarcoding differentiates between micro-habitats within the rocky intertidal (Shea & Boehm, 2024). Version 1.5. United States Geological Survey. Occurrence dataset. https://doi.org/10.15468/33artc accessed via GBIF.org on 2025-05-07.
We flattened the data into a single table with the columns in alphabetical order. See if you can:
- map the column headers to DwC and the DNA extension
- split the data into two files: an occurrence table (Darwin Core terms) and a DNA table (DNA extension terms)
Download the file here: https://sunray1.github.io/2025-05-09-GBIF-NA-DNAPublishing/files/DNA_example_2_for_IPT.csv.zip
Key Points
No two datasets are the same, you should think about your dataset before you start mapping
There are hundreds of Darwin Core terms, lean on the guides when you’re mapping your dataset